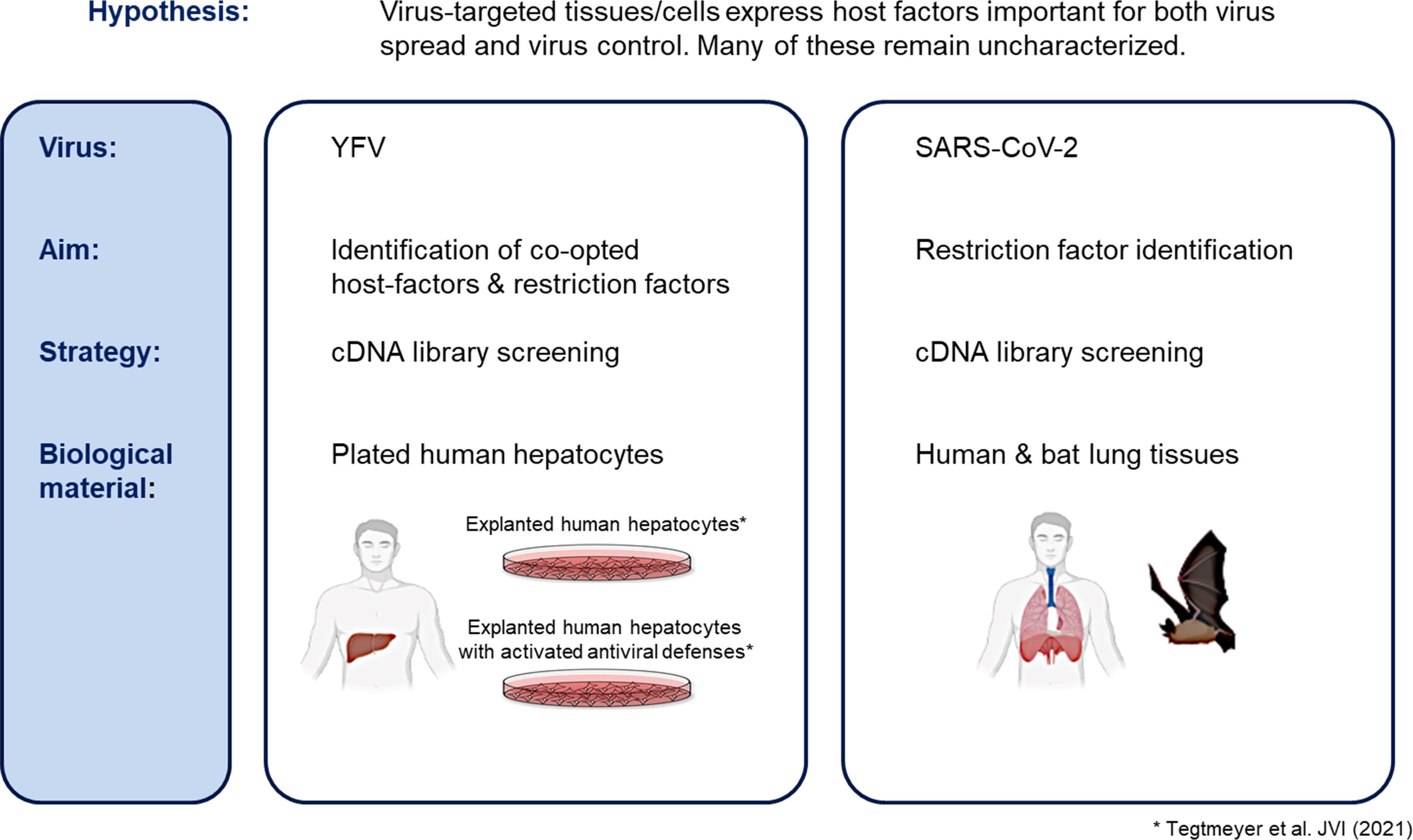
Viral and host-cell proteins establish a range of molecular interactions during infection. This virus-hostinteractome is unique for each virus and host species. Many host-cell factors are hijacked during infection and contribute to viral spread. At the same time, antiviral factors are induced which target different viral proteins and suppress virus production. This project aims to identify novel factors which either promote or suppress the spread of two pathogenic viruses: yellow fever virus (YFV) and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). To achieve this, we use mRNAs extracted from primary tissues targeted by these viruses: human hepatocytes and human/bat lungs. These mRNA pools are cloned into lentiviral vectors to create complementary DNA (cDNA) libraries, which encode a tissue-specific pool of proteins. These cDNA libraries are then transduced into selected cell-lines, and the pool of ectopically introduced genes express mRNAs and proteins, which can be quantified by total RNA-sequencing. Transduced cell-lines can then be used for ‘loss-of-function’ or ‘gain-of-function’ screening, performed with live YFV and SARS-CoV-2. After screening, surviving cells are assessed to determine whether an increase or reduction of virus infection rates is observed. If so, the factors responsible can be identified by RNA sequencing, by comparing frequencies of all library delivered gene products in the cell population before and after screening. Ultimately this project aims to identify human factors that facilitate virus spread and could represent therapeutic targets. Additionally, we aim to identify human and bat factors that facilitate tolerance to infection-induced pathology.
Contact details:
Dr. Richard J P Brown
Group Leader, Department of Veterinary Medicine
Paul-Ehrlich-Institut
Federal Institute for Vaccines and Biomedicines
Paul-Ehrlich-Str. 51-59
63225 Langen
Phone +49 6103 77 7441
E-Mail Richard.Brown@pei.de
Copyright for figure: Images created with Biorender.